Description
It has long been a worldwide problem to precisely identify leucine and isoleucine, not to mention 100% accuracy. Creative Biolabs, the leader in De Novo antibody sequencing filed, never forgets to push the boundaries of science. They have successfully differentiated the sequences of leucine and isoleucine with 100% accuracy after overcoming challenges and difficulties.The amino acid sequence and specific post-translational modifications determine the specificities and effectiveness of an antibody drug, which confirms the indispensable role of sequence analysis during monoclonal antibodies development. The traditional way of DNA sequencing cannot fulfill the analytics requirements of some pre-clinical antibody drugs derived from an immunized host or a hybridization, whose cDNA sequences are unavailable in the current database, thereby antibody de novo sequencing at protein-level on the scene occurs.
Based on analyzing the regular breakage of peptide fragments, antibody de novo sequencing is a protein-level analysis independent of any existing protein database information. When colliding the specific fracture mode with inert gas, corresponding amino acid sequence information and post-translational modification can be calculated. Half a century ago, scientists used a technique named Edman degradation, which is still applied today in some short peptides sequencing, to gradually degrade protein polypeptide from N-terminus to C-terminus. Each degraded amino acid residue will be identified by detecting the ultraviolet absorbance during the degradation process. However, apart from the error accumulated during this cyclical process, this chemical method cannot satisfy the requirements for high-purity samples and be used for complex analysis procedures.
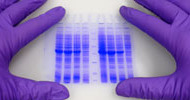
In the last two decades, mass spectrometry, another prominent technology to sequence long peptides has been gradually developed. The protein samples need to be digested before analyzed by mass spectrometry. To maximally obtain the fragment information, the data collection process is complicated by the combination of various techniques, including the high-resolution and high-quality precision mass spectrometry, multiple enzyme digestion, and diverse mass spectrometry fragmentation methods. The spectrums are generated by the molecular weight difference, hence determining the amino acid residues. The final monoclonal antibody amino acid sequence information is then obtained after analysis and splicing with the antibody de novo sequencing software.
Soft ionization charges the isolated peptide fragments before scanned by mass spectrometry to form the MS spectrum. These screened peptide ions will then enter the collision chamber that contains inert gases, such as He, Ar, Xe and CH4, to induce regular covalent bond cleavage of the peptide, generating daughter ions to create the MS-MS spectrum in the mass spectrometry.
At present, the commonly used cleavage methods of data acquisition in mass spectrometry are collision-induced activation dissociation (CID) and high-energy collision dissociation (HCD) suppled with electron transfer dissociation (ETD). Since the MS-MS spectrum produced by HCD has higher quality and theoretical suitability. HCD can be applied in secondary cleavage to produce corresponding b and y ions. Besides functions in better determining amino acid sequences by generating c, z ions complementary to b, y ions, ETD can accurately confirm the post-translational modification site of the protein via retaining post-translational modification groups that are easily lost in the side chain. Besides, characteristics of electron transferring render the ability for ETD to realize the cleavage of long peptide fragments with a high amount of electric charges and low mass-to-charge ratio.
Depending on the peptide cleavage sites, ions will gain different features. For instance, the ions generated near the N-terminus are type a, b, and c, while near the C-terminus are x, y, z. B-type ions are special in identifying amino acids because the cleavage bond is between two amino acid residues. The weight value difference between the two adjacent-b ions equals to that of an R group, depending on which amino acid type will be confirmed by comparing the value of the R group.
Unfortunately, not every amino acid can be identified easily. Leucine and isoleucine, for example, randomly distributed in an antibody, have identical weight and similar chemical properties. Researchers normally estimate their location by arrangement when determining the sequence and function of antibodies, which is time-consuming and uneconomical. Recently, scientists at Creative Biolabs overcame a technical challenge by accurately differentiating the locations of leucine and isoleucine, enabling antibody sequencing for better functional analysis, validation and subsequent applications.
De novo antibody sequencing presents significant application values, such as the recovery of lost information of gene and disease-specific monitoring, selection of isotype recombinant chimeric antibodies, and proof of bispecific antibody optimal target combination. More importantly, it can be applied to develop biosimilars by determining sequences and innovative sequence variants from biopharmaceuticals.
One error in amino acid sequencing may lead to large changes in antibody function during sequencing. The tremendous diversity is created by gene recombination and hypermutation introduced by B cells, a method to improve antibody affinity, making de novo protein sequencing of monoclonal antibodies an enormous challenge. In addition to overcoming common difficulties and challenges such as sequencing of V, J and C gene segments or CDR3 domain, Creative Biolabs offers unequivocal sequencing for variable domains using mass spectrometry to identify leucine and isoleucine with 100% accuracy.